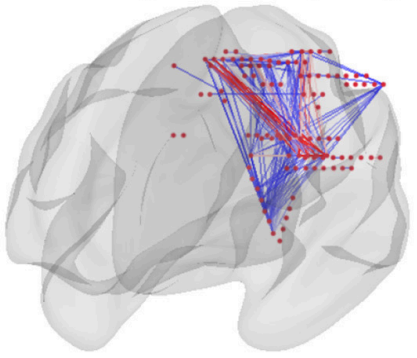
Research
Topic and highlights of my research
Postdoc researcher in computational neuroscience


Topic and highlights of my research
List of publications and review work for journals

Courses and diplomas

Open-source Python softwares